PhD defense of Freerk Venhuizen from the Eye Lab
December 10, 2019
Sanne van Velzen receives Best Student Paper award at SPIE MI 2020
February 21, 2020Our paper ‘A Deep Learning Model for Segmentation of Geographic Atrophy to Study its Long-Term Natural History’, by Liefers et al, has been published in Ophthalmology. In this paper, we develop and validate a deep learning model for the segmentation of geographic atrophy (GA) in color fundus images. With this model, we were able to fully automatically segment GA in 3589 images in an independent data set (from the AREDS). We used the automatically segmented GA area to study the variability in growth rate: the atrophic area grows much faster in some eyes compared to other eyes. We found that part of this variability can be explained by the presentation at baseline: features describing the shape, size and distribution of the baseline GA area predict future growth rate. Moreover, we found that most eyes initially exhibit quadratic growth of the GA area. However, the growth rate levels off if the area increases further.
Currently, there is no cure for GA. However, several potential therapies are being investigated in clinical trials. The goal of these therapies is to slow down the growth of GA. To validate the effectiveness of these therapies, it is important to understand the natural history of GA. With this work, we contribute to that understanding.
One technical challenge for getting the model to work was the large variability in image quality, with data form different camera’s, containing both digital and digitized (scanned analogue) images. The resulting model is therefore quite robust to these variations, and can now be used in many other data sets. We are particularly excited about the opportunity to study the relationship between imaging parameters and GA growth rate, as well as in the context of genetics, demographic information and life-style.
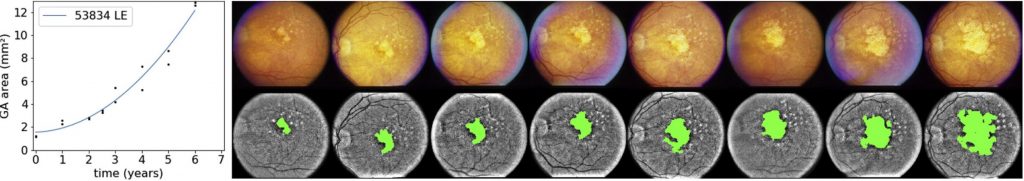
Segmentation of GA to study its long-term natural history.